Cloning Vectors: Essential Features
by Pallabi Roy Chakravarty, Ph.D.

by Pallabi Roy Chakravarty, Ph.D.
A “vector,” in simple terms, is an agent for transmission. For example, mosquitoes are vectors for malaria; western blacklegged ticks are vectors that transmit Lyme disease to humans. Think of them as the transport vehicle for its passenger.
Let us understand what vectors are in the context of cloning procedures for molecular biology and biotechnology purposes.
In this case, vectors are the DNA molecules used as tools to transfer a foreign DNA fragment of interest (FoI) to the host cell. This ensures that the FoI is stably replicated there.
Additionally, in some cases, the vectors are suitably designed to facilitate adequate expression (transcription and translation) of the FoI in the host cell. This type of vector is called an expression vector.
In this article we will explore the four essential features that are required for cloning vectors, including their relevance and impact on the cloning process. A schematic representation of a vector molecule including its essential features is depicted in figure 1.
Figure 1. A vector, including the essential sequence elements in its backbone.
Article Contents:
1. Origin of replication (Ori) in the host cell
Significance of origin of replication (ori) in molecular cloning
Why are vectors with a stringent control ori low copy number?
Mechanism of relaxed control of vector replication
2. Multiple Cloning Site (MCS)
4. Sequence elements for expression of cloned DNA fragment
4B. Sequence elements for translation
A vector must have the necessary nucleic acid sequences to ensure that it can be replicated within the host cell.
Ori is the sequence on the vector where its replication is initiated using the host cell’s replication machinery (DNA polymerase and other enzymes).
Generally, ori sequences are rich in A and T, and low in G and C nucleotides. A-T pairs have two hydrogen bonds while G-C pairs have three. Thus, AT rich sequences at the ori region separate easily, making the DNA single-stranded, which is important for the DNA replication to occur smoothly.
A vector replicon comprises the ori and all other sequence elements that regulate its replication.
A schematic diagram of the replicon region of a vector is shown in figure 2.

Figure 2. Representation of the replicon, including the ori.
The origin of replication controls the copy number of the vector.
Copy number is the number of copies of a vector molecule that is maintained in the host cell. This is inherently tied to the replication efficiency of the vector.
There are two mechanisms that control replication from a vector’s ori: stringent and relaxed.
Generally, vectors with relaxed replication control have high copy number, while those with stringent control have a low copy number.
With the relaxed mechanism, replication of the vector can be initiated independent of the host cell’s replication machinery. Only the elongation and termination steps require host machinery.
Stringent control is characterized by needing the host’s replication machinery for the initiation of vector replication. This characteristic is the basic difference between the relaxed and stringent regulatory mechanisms.
In stringent regulation of replication, host cell proteins such as DnaA and integration host factor (IHF) are required for initiation of vector replication.
Thus, replication is directly controlled by concentration of replication-initiation proteins encoded by the host.
Protein synthesis is a metabolically expensive process for the host cells. For this reason, the host cell tightly controls protein synthesis, including those required for replication to prevent unnecessary metabolic load on itself.
Consequently, the replication of these vectors, and therefore their copy numbers, are limited due to the limited availability of host replication-initiation proteins.
When it comes to relaxed vectors, initiation of replication is not dependent on the host cell’s metabolic status. Rather, the inhibition of vector replication is facilitated by vector encoded RNA transcripts, whose concentration increases with vector concentration in the host cell.
Consequently, vector replication is not inhibited until a high number of vector molecules are produced in the cell. This explains the high copy number of such vectors.
As an example, let us consider the case of ColE1 plasmids. Here, an antisense RNA encoded by the vector stalls the replication process by binding to another RNA that is required as a primer to initiate plasmid replication.
A schematic representation of this method of inhibition is shown in figure 3.

Figure 3. Regulation of plasmid replication by self-encoded antisense RNA
a) Low plasmid concentration in the host cell. The plasmid encodes a non-coding RNA named RNAII. This RNA is critical in plasmid replication. It gets modified by the enzyme RNaseH and subsequently binds to the leading strand of the ori region, like a primer. This enables DNA polymerase to drive replication (DNA polymerases require primer-template double-stranded junction to function).
b) High plasmid concentration in the host cell. Another non-coding RNA, RNAI is transcribed by the plasmid. RNAI is antisense to RNAII and is able to base-pair with RNAII. This double-stranded RNA complex is further stabilized by another plasmid-encoded product, the protein Rpo (orange box). This RNAII-RNAI duplex is now inaccessible to RNaseH. Ultimately, this leads to the inability of singe-stranded RNAII, adequately processed by RNaseH, to prime to the leading strand at the ori. This results in turning off plasmid replication. Note, expression of both RNAI and Rpo are positively dependent on plasmid number in the host cell.
Let us illustrate another case of the relaxed control of plasmid replication. RepA is a plasmid encoded replication-initiation protein required for vector replication. An antisense RNA transcribed from the vector binds to the repA mRNA thereby preventing RepA protein translation. This ultimately blocks the plasmid’s replication. However, this antisense RNA concentration is dependent on the plasmid’s concentration. Thus, until a high number of plasmids is present in the cell, this method of replication inhibition does not work and leads to such plasmids having high copy number.

Figure 4. RepA is a plasmid encoded protein that binds to the plasmid replicon and facilitates initiation of replication. As shown in scenario (a), when the plasmid concentration in the cell is low, the RepA protein is synthesized and it drives replication. When plasmid concentration is high (by adequate replication), as depicted in (b), a non-coding RNA copA is transcribed by the plasmid, which binds to the repA mRNA and blocks its translation. Thus, with the RepA protein production being inhibited by this mechanism, plasmid replication is ultimately turned off. The concentration of copA RNA is dependent on the levels of the plasmid in the cell. Consequently, this method of plasmid replication negative regulation is effective when there is a high number of plasmids inside the host cell.
Another area of ori significance has to do with vector incompatibility. When two different vectors cannot be maintained in one host cell, they are called incompatible. The two vectors may be of the same kind or of a different kind (for instance, one vector and one phagemid
The replicon elements, most prominently the ori, plays a determining role in vector compatibility.
As mentioned above, in relaxed controlled vectors, negative regulation systems encoded by the vector itself modulate replication. In the case of incompatible vectors, the ori is the same. Thus, the negative regulators, for example, the antisense RNAs described above, cannot distinguish between the different vectors present within the same cell. Consequently, the replication regulation is erroneous.
However, when vectors are compatible, the negative regulator for each vector is different, and thus the corresponding negative regulator can correctly identify and regulate the vector under its own regulation.
There is yet another reason for plasmid-plasmid incompatibility. This is related to how two plasmids are partitioned to daughter cells upon cell division.
The next essential feature of a plasmid cloning vector is the multiple cloning site (MCS). Appropriate sequences are required on the vector molecule that facilitate easy incorporation of the fragment of insert into it. The MCS serves this purpose. A multiple cloning site is a small segment of DNA on the vector that contains multiple restriction enzyme cut sites. These sites are important for restriction enzyme recognition and action.
Most commonly, the MCS has multiple restriction endonuclease cleavage sites. For cloning of the FoI into the vector, the FoI and the vector are digested with the same restriction endonucleases.
This produces “sticky ends” in the vector and the FoI intended to be inserted (also called the insert) within it.
Such a complementarily digested vector-insert pair is thereafter joined together using the enzyme DNA ligase.
For details of vector-insert restriction endonuclease-digestion and ligation for molecular cloning, please refer to this article.
The presence or absence of the vector within the host cell needs confirmed. Furthermore, it is important that such a confirmation of vector presence in the hose cells can be done easily using a readily distinguishable visual phenotype.
For this purpose, the vector encodes at least one gene that confers a distinguishing phenotype to the host cells. Host cells possessing this gene can easily be compared to those lacking the gene.
Frequently, such marker genes in the vector that facilitate selection (also called selectable markers) are antibiotic resistance cassettes.
When host cells are plated on media containing the selection pressure (the corresponding antibiotic), only the ones having the vector will grow (positive selection) while the cells without the vector die. A representation of the role of a selectable marker is depicted below in figure 5.

Figure 5. Illustrates the concept of selection using media containing selection pressure. Cells that did not take up the plasmid will not grow when plated. However, host cells that did take up the desired plasmid will have colonies growing on the plate.
For details about how selectable markers within vectors are used in molecular cloning, please refer to this article.
While the above features are necessary for all vectors, the following described below are characteristic elements of expression vectors:
For transcription of the cloned FoI, a promoter sequence is required on the vector that utilizes the host cell’s transcription machinery to drive transcription of the FoI. The architecture of the vector backbone is designed so that the FoI can be cloned downstream of the promoter.
Promoters comprise a stretch of DNA sequences where RNA polymerase binds and initiates transcription.
A representation of a vector’s promoter and other sequence elements for the expression of DNA fragments cloned in the MCS is shown in figure 6.
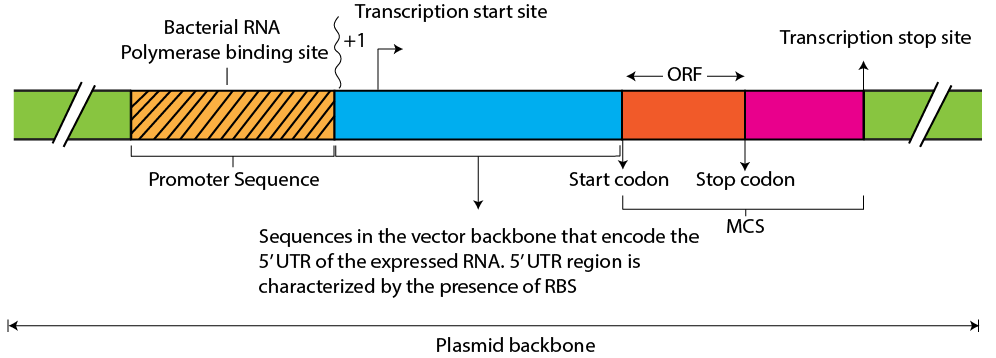
Figure 6. The promoter region and immediate downstream sequences, including a hypothetical ORF cloned in the MCS
Based on the need of the experiment, the promoter can be chosen or engineered to drive constitutive expression of the FoI cloned downstream.
Constitutive promoters are unregulated promoters that allow continuous transcription of its downstream gene.
Alternatively, researchers can choose conditional promoters for tight expression control.
Conditional promoters drive transcription only under certain environmental stimuli such as at a specific temperature or in the presence of a certain sugar, such as lactose or its analog.
If it is intended to express the protein product of the FoI cloned on the vector, then suitable genetic sequence elements are required in the vector backbone to ensure proper translation of the FoI mRNA. Minimal requirements are:
Depending on the host cell, these essential sites need to be optimally designed. For example, if the host cell is a bacterium, the vector needs to have the Shine Dalgarno sequence for the bacterial ribosomal machinery to effectively drive translation of the FoI mRNA product.
Further, depending on the need of the experiment, the vectors may have appropriate signal sequences that facilitate the addition of specific amino acid sequences into the final protein product of the cloned FoI.
For example, expression vectors can be chosen so that a stretch of six histidine residues are appended to the FoI protein. Such a His-tagged protein can be purified using a Nickel-NTA column.
In other cases, appropriate amino acids can be appended to the FoI protein using sequences of the vector, which may facilitate partitioning of the cloned protein to the host cell membrane or transmembrane region.

In the microscopic world, different species of bacteria and fungi are engaged in a never-ending turf war, fighting to defend their territory and to advance...

Regular agarose beads are typically used in a column format to purify molecules and perform immunoprecipitation or pull-down experiments. Magnetic agarose beads are an excellent...

Do you ever feel like you’re bouncing through life, squeezing through the never-ending to-do items on your list? If so, then you have a good...

Do you use an AirTag®, Tile®, or other Bluetooth® tracker to keep tabs on anything? When people use these types of tracking devices, it is...
