How to Choose Agarose Resins for Protein Purification
by Tyasning Kroemer, Ph.D.

by Tyasning Kroemer, Ph.D.
Purifying a target protein is an important step before investigating its structure and function. To purify a target protein from a mixture of substances, column chromatography is a useful technique due to its convenience and cost. This article provides a guide to help you choose the right agarose resins for protein purification using column chromatography.
In this article:
What is Column Chromatography?
Column Chromatography Preparation
What are Agarose Resins (Agarose Beads)?
What Features are Important to Consider before Choosing Agarose Resins?
Column chromatography is a common technique, used for research in protein biochemistry. This method is useful to purify a protein based on their features, such as its size or specific binding affinity to a ligand.
For example, to separate small molecules from unwanted large molecules in your sample, choose size exclusion chromatography by using a column and a porous matrix.

Otherwise,
to separate a protein using its specific binding affinity to a ligand, use agarose
beads cross-linked to a ligand.

To learn more about how column chromatography works, read GoldBio article below:
How Column Chromatography Works to Separate Proteins
To perform column chromatography, in addition to your sample, you will need a column, a matrix, a solvent to dissolve your sample, a washing buffer, and an elution buffer.
After gathering your supplies, you’ll need to pack the column with a matrix, such as agarose beads or resins. Keep in mind that you must avoid getting bubbles when packing your column.
To learn how to pack your column, watch this video below:
Agarose resins or agarose beads are matrix, which are composed of agarose and used in column chromatography.
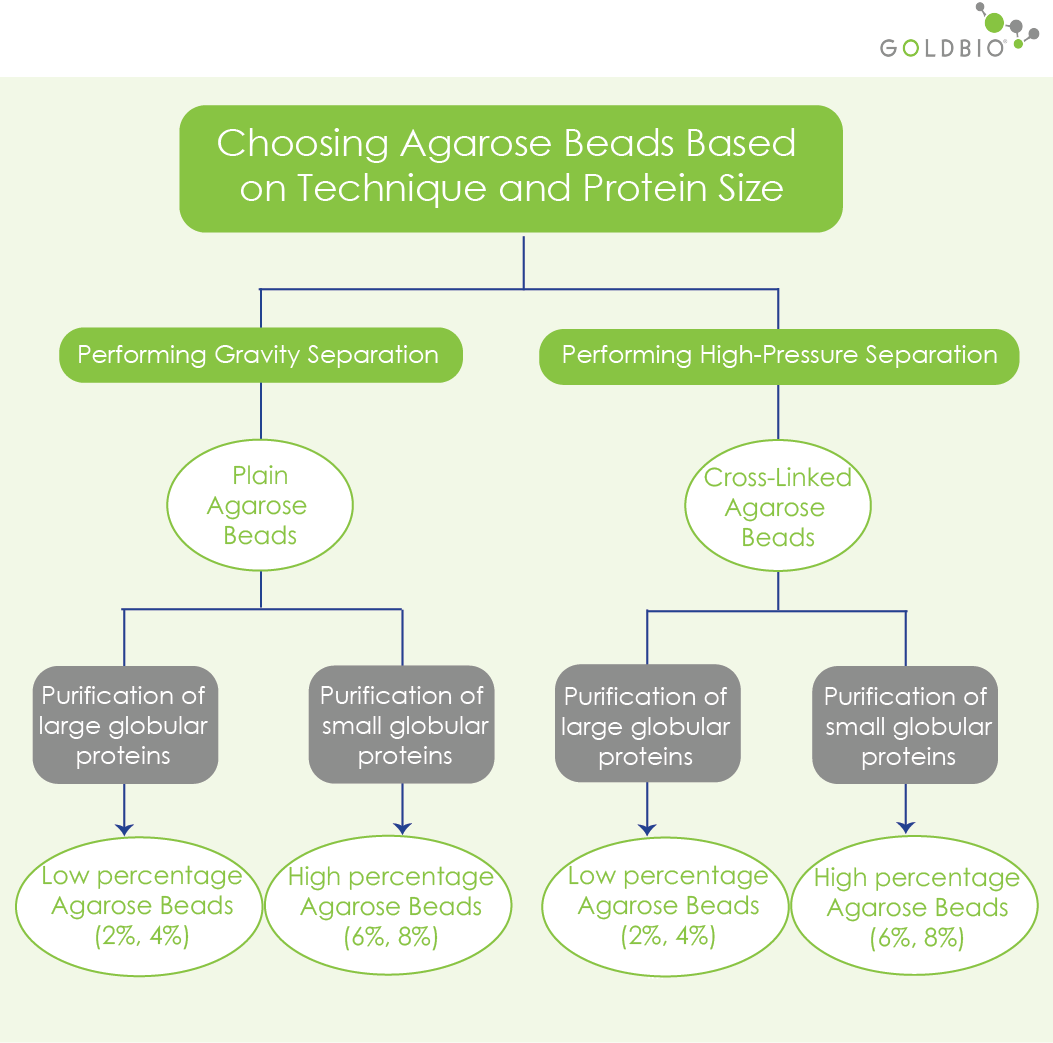
To separate proteins by their size, choose plain or cross-linked agarose beads.
Plain agarose beads are sufficient when using gravity techniques, but not for use in high-pressure separation techniques. Gravity techniques use gravity to move the mixture of your sample and solvent (or mobile phase), whereas high-pressure techniques uses compressed gas to push the mobile phase in the column.
Cross-linked beads are physically and chemically more stable than the plain agarose beads. Cross-linked beads are high-density agarose beads, treated with a particular reagent to produce more cross-linked polysaccharide chains, to add the strength and stability to the matrix.
Remember, when performing chromatography, you’re pouring a solvent through the column. This can cause pressure that would otherwise collapse the matrix. Therefore, physical stability in this situation is critical. In addition, the matrix should not react with any of the solutions during the purification.
For both plain and cross-linked agarose resins, there are different percentages of agarose resins: 2%, 4%, 6% or 8%, allowing better separation of molecules based on their size. The pore sizes are smaller in the higher percentage of agarose beads, so it’s useful to separate small globular proteins. Likewise, the lower percentage of agarose resins is better to separate larger globular proteins.
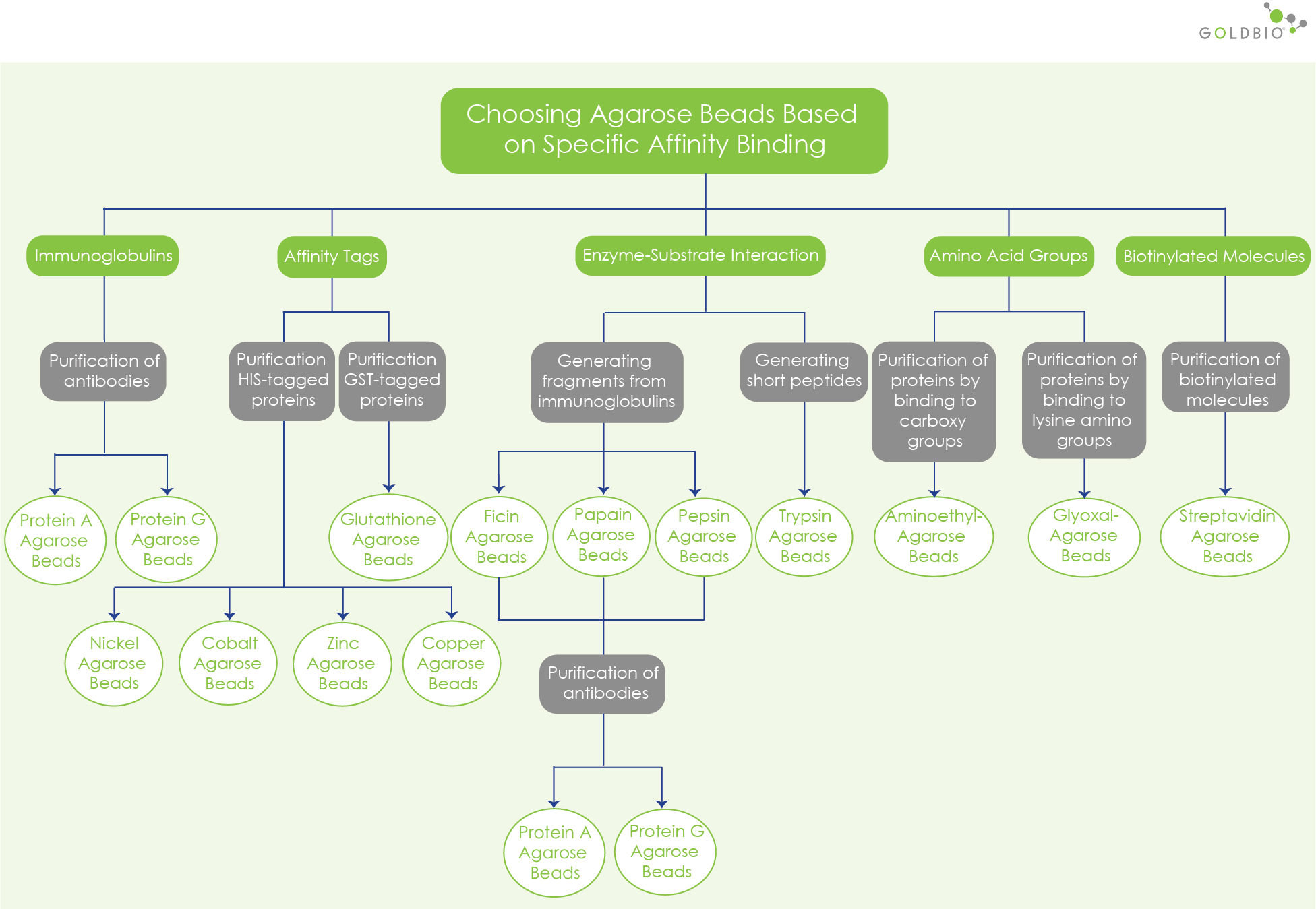
Another consideration when choosing your beads is specific binding affinity. It is another technique to purify your protein of interest by binding it to a ligand.
Some examples of protein features and structures used for affinity chromatography:
If your protein is comprised of carboxy group amino acids, aminoethyl-agarose beads allow covalent binding of the amino groups with carboxy groups. However, choose glyoxal-agarose beads, if your protein is comprised of lysine amino group.
Affinity tags are short peptide sequences linked to your recombinant protein. These short peptide sequences often encode for a protein used for purification, labeling, detection, or visualization. During protein purification, an affinity tag of a protein binds to a ligand linked to the agarose resin.
Two common affinity tags used for protein purification:
The GST-tag is a short protein encoding an enzyme, gluthathione S-transferase. This enzyme binds to a glutathione. For a protein with the GST tag, choose glutathione agarose resin. Glutathione is a highly efficient affinity purification tool because its affinity for GST is in the submillimolar range.
The His-tagged proteins are proteins with a short sequence containing a string of histidine residues. The histidine tags have affinity for metal ions.

Purify His-tagged proteins by choosing agarose beads containing metal ions to perform immobilized metal affinity chromatography (IMAC). IMAC is a method to purify a protein with an affinity for metal ions, such as Co2+, Ni2+, Cu2+, or Zn2+, immobilized on a matrix (Yang et al., 1998).
NTA vs. IDA Agarose Beads
The metal ions are fixed on cross-linked agarose beads by a chelating ligand, nitilotriacetic acid (NTA) or iminodiacetic acid (IDA). NTA is more stable than IDA because it contains four chelating sites. IDA only has three chelating sites.
GoldBio offers products for this purpose, such as: nickel agarose beads and cobalt agarose beads. A high-density resin helps increase the affinity of the resin, whereas a low density increases the specificity of the resin.
To learn more about His-Tag affinity for metal ions, read GoldBio article below:
His-Tag Metal Affinity Cations: What’s the difference again?
Bacterial proteins serve as useful ligands for purifying polyclonal or monoclonal immunoglobulin G (IgG). These bacterial proteins include protein A and protein G. Both protein A and protein G , can bind to human and mammalian IgG antibodies, so they are useful as a ligand. However, protein A and protein G may have different affinities for IgG subclasses from different species.
Protein A and protein G bind to the Fc region of the immunoglobulins. Protein A binds to IgGs from different mammalian species, whereas protein G is effective for isolating, purifying immunoglobulins from cell culture media or biological fluids.

For antibody purification and immunoprecipitation, you can choose GoldBio protein A agarose resin or protein G agarose resin.
The specific interaction between enzymes and their substrates provides a good tool for cleaving your molecule into fragments. This approach to enhance the performance of antibodies uses enzymes immobilized on agarose beads.
For purifying your antibody, you can then use a column with protein A or a column with G agarose beads to collect your specific fragments.
Some examples of these resins are:



A bacterial protein called streptavidin interacts non-covalently to a small molecule biotin (vitamin H/vitamin B7), making this protein an important tool for many molecular biology applications. As an example, to purify biotinylated antibodies, streptavidin agarose resin is useful for gravity flow columns, spin columns, and fast protein liquid chromatography.
Find GoldBio products below:
Berg, J. M., Tymoczko, J. L., & Lubert Stryer. (2013). The Purification of Proteins Is an Essential First Step in Understanding Their Function. Nih.Gov; W H Freeman. https://www.ncbi.nlm.nih.gov/books/NBK22410/
Block, H., Maertens, B., Spriestersbach, A., Brinker, N., Kubicek, J., Fabis, R., Labahn, J., & Schäfer, F. (2009, January 1). Chapter 27 Immobilized-Metal Affinity Chromatography (IMAC): A Review (R. R. Burgess & M. P. Deutscher, Eds.). ScienceDirect; Academic Press. https://www.sciencedirect.com/science/article/pii/...
Böing, A. N., van der Pol, E., Grootemaat, A. E., Coumans, F. A. W., Sturk, A., & Nieuwland, R. (2014). Single-step isolation of extracellular vesicles by size-exclusion chromatography. Journal of Extracellular Vesicles, 3(1), 23430. https://doi.org/10.3402/jev.v3.23430.
Chivers, C. E., Koner, A. L., Lowe, E. D., & Howarth, M. (2011). How the biotin–streptavidin interaction was made even stronger: investigation via crystallography and a chimaeric tetramer. Biochemical Journal, 435(Pt 1), 55–63. https://doi.org/10.1042/BJ20101593.
Coskun, O. (2016). Separation Techniques: CHROMATOGRAPHY. Northern Clinics of Istanbul. https://doi.org/10.14744/nci.2016.32757.
Enzyme Immobilization on Polymeric Resins. (2008, October 1). GEN - Genetic Engineering and Biotechnology News. https://www.genengnews.com/magazine/99/enzyme-immo...
Gustavsson, P.-E., & Larsson, P.-O. (1996). Superporous agarose, a new material for chromatography. Journal of Chromatography A, 734(2), 231–240. https://doi.org/10.1016/0021-9673(95)01304-0.
Mohamad, N. R., Marzuki, N. H. C., Buang, N. A., Huyop, F., & Wahab, R. A. (2015). An overview of technologies for immobilization of enzymes and surface analysis techniques for immobilized enzymes. Biotechnology & Biotechnological Equipment, 29(2), 205–220. https://doi.org/10.1080/13102818.2015.1008192.
Stellwagen, E. (1990, January 1). [25] Gel filtration (M. P. Deutscher, Ed.). ScienceDirect; Academic Press. https://www.sciencedirect.com/science/article/pii/007668799082027Y.
Yang, Q., Tomlinson, A. J., & Naylor, S. (1998, January 1). Chapter 3 - Sample Preparation (Z. Deyl, I. Mikšik, F. Tagliaro, & E. Tesařová, Eds.). ScienceDirect; Elsevier. https://www.sciencedirect.com/science/article/abs/pii/S0301477008603012.

Labeling antibodies with biotin or a fluorophore enables scientists to detect, track, and quantify that antibody and the molecules that it binds to. We cover...

Covalently conjugating a small molecule to an antibody’s surface is a process called antibody “labeling.” Labeling antibodies with small molecules such as biotin or fluorophores...

DNA gel electrophoresis is one of the most widely used analytical techniques in molecular biology, providing a simple and reliable way to separate DNA fragments...

Do you have a favorite restaurant that you love because you know exactly how great the experience is going to be? There are probably a...
